Vurtigo Features include:
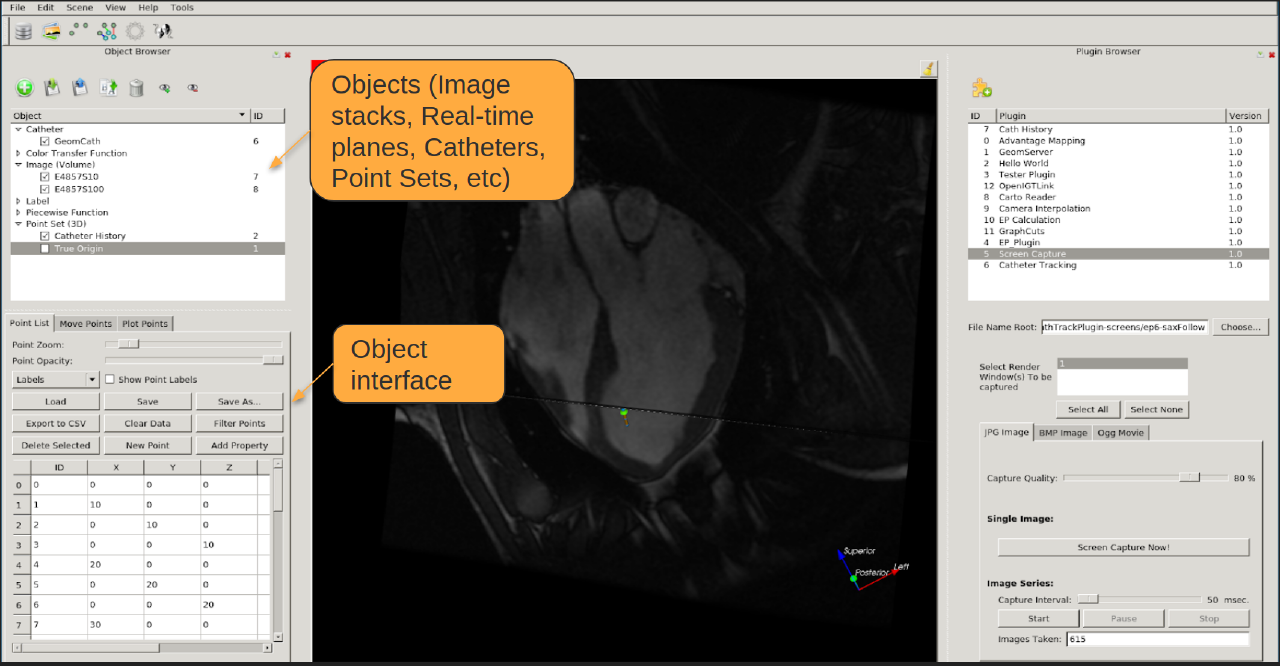
Extensible Plugin Architecture The Vurtigo core provides support for rendering and controlling objects. Plugins provide the main features, input/output, etc. Plugins can be licensed separately (SDK to be released soon)
3D/4D Roadmaps Load and navigate multiple DICOM datasets; supports cine datasets
Multiple Modality Datasets MR, CT (DICOM images with geometry information), JPEG/PNG
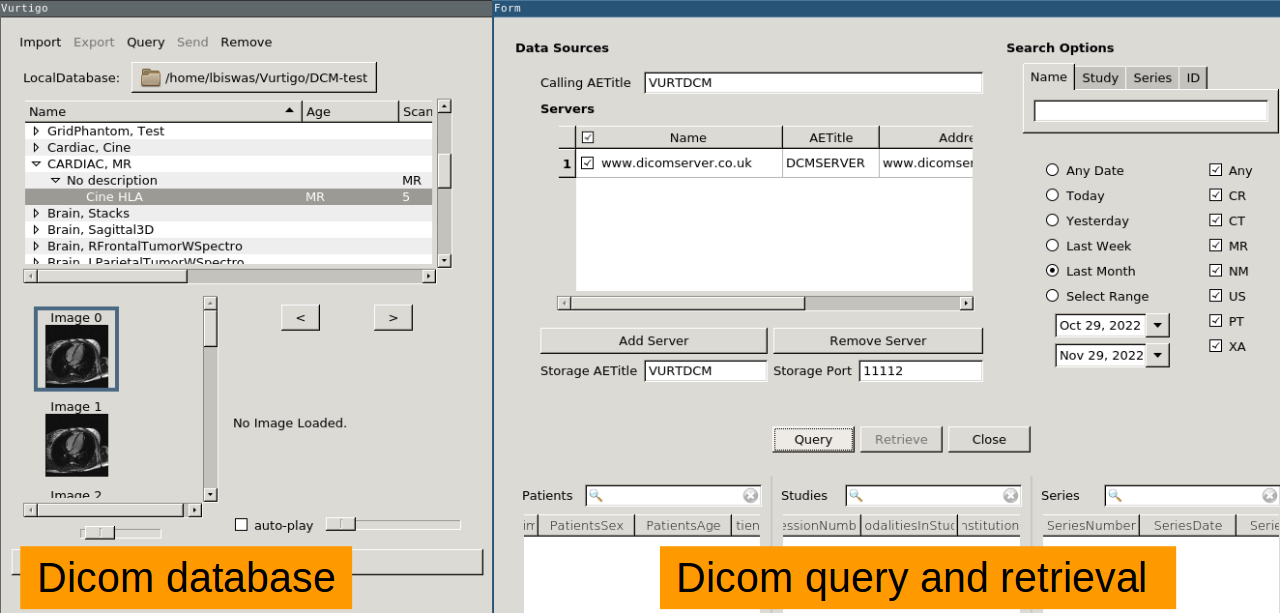
DICOM Database Storage of DICOM images. Query and retrieve images from remote DICOM nodes
Volume Rendering Configurable rendering modes and properties, GPU acceleration
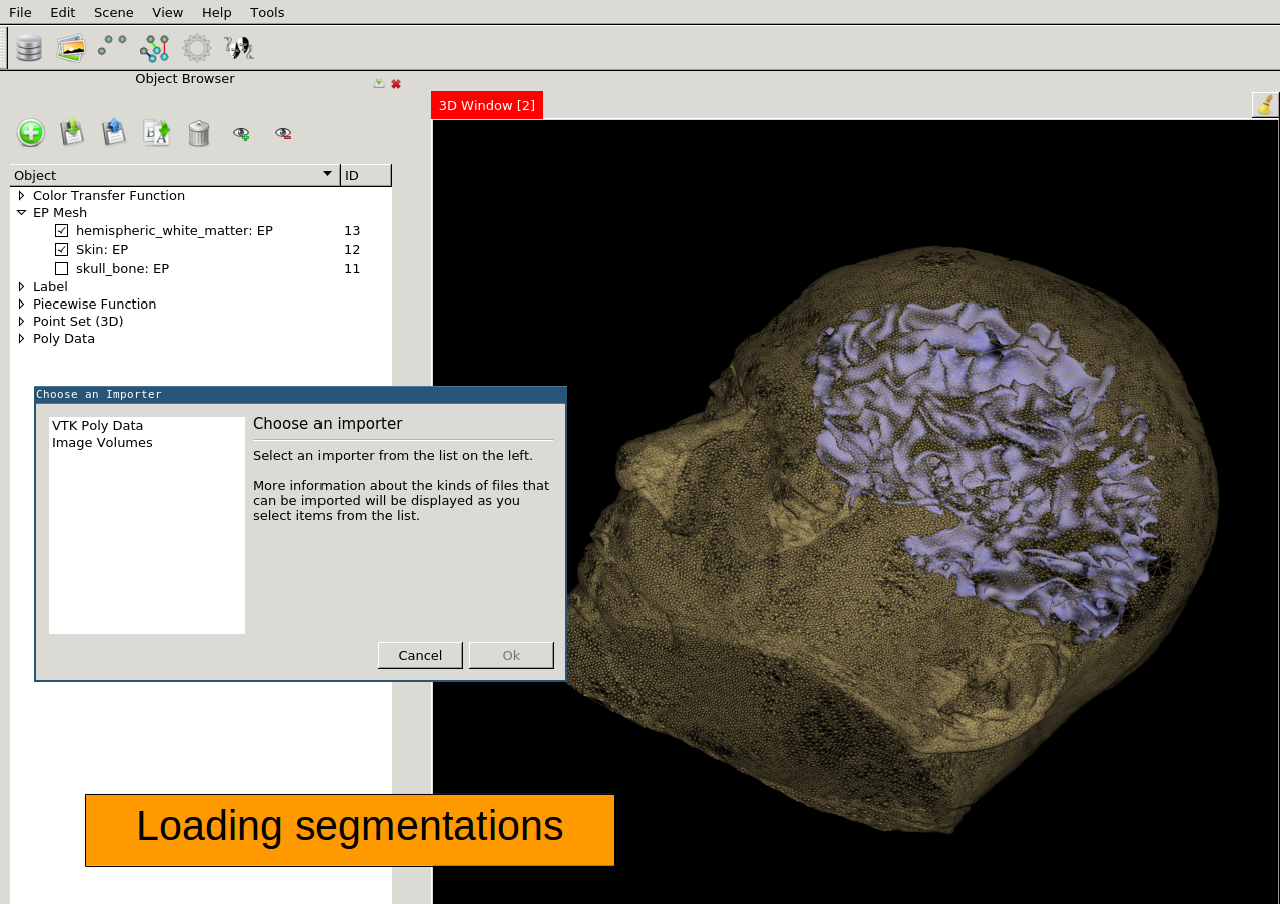
Surface Rendering Can be used to display segmented datasets
Mesh Painting Paint a surface using data from points
Configurable Visualization Layout 3D Only, 2D + 3D views, 2D only, multiple 3D windows
3D Points Create, manipulate and filter (based on configurable properties) 3D point clouds
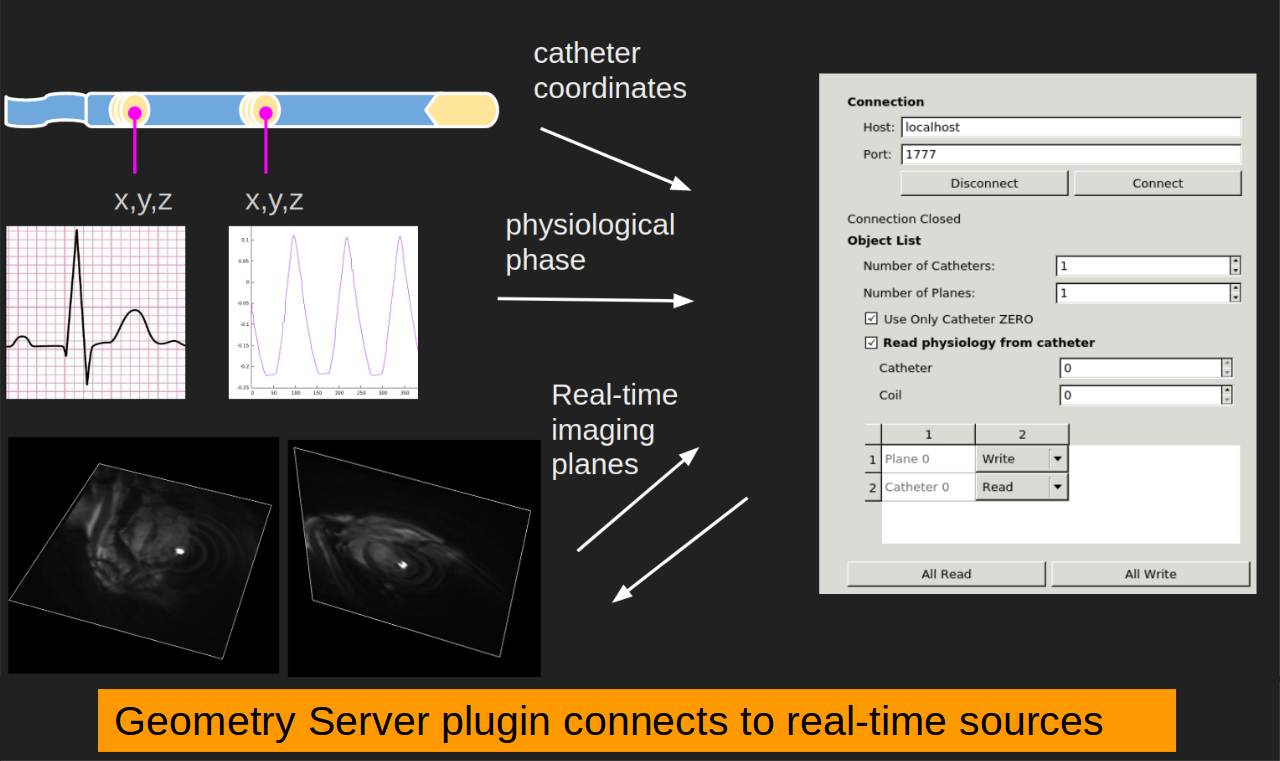
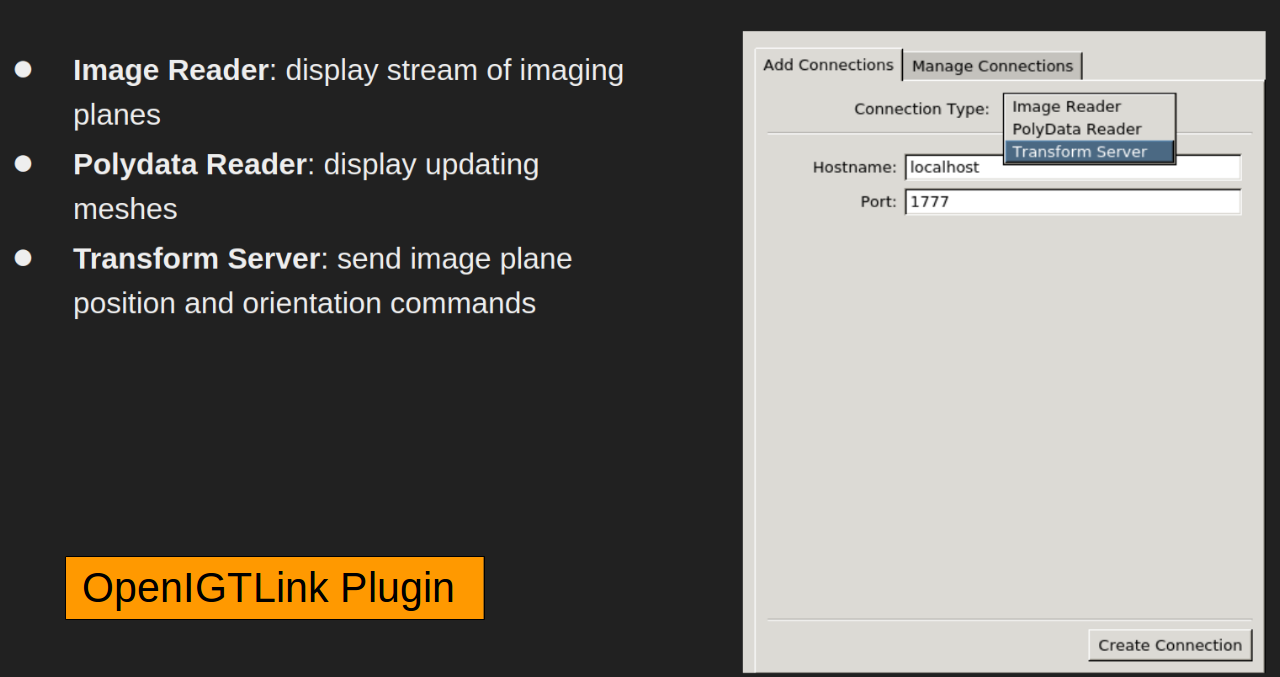
Scanner Communication Can connect to real-time scan information via the Geometry Server (included) or OpenIGTLink. The Geometry Server can work with compatible scanner console software such as Vista.ai’s RTHawk Research (not included)
Real-time Imaging Display and control multiple interactive real-time imaging planes within the 3D view
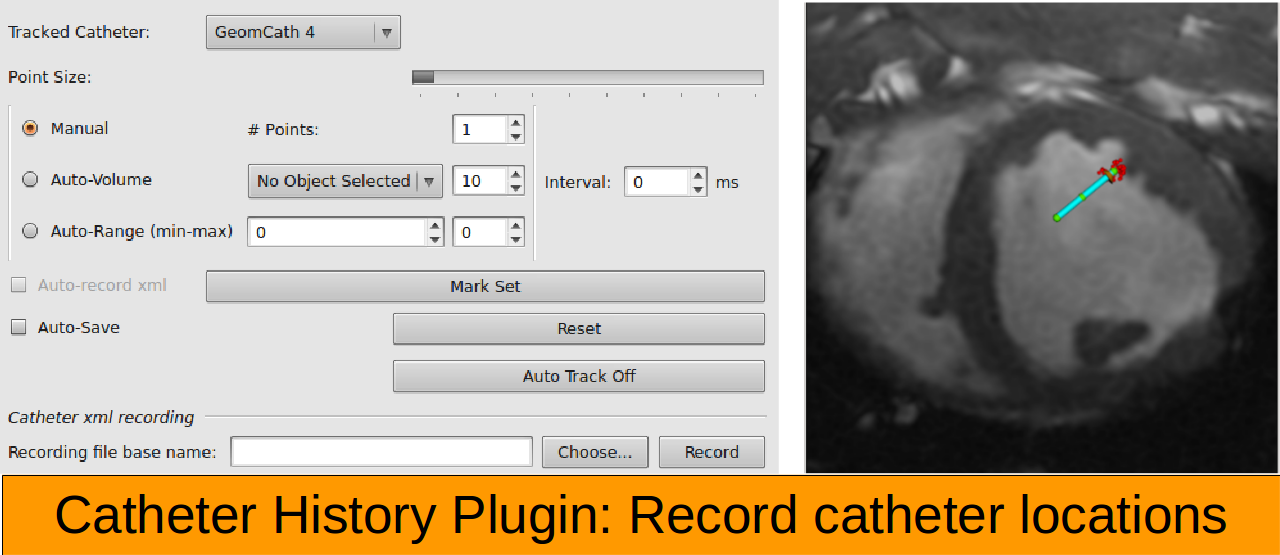
Catheter Visualization Display multiple actively tracked catheters in 3D view
Catheter Following Update real-time imaging planes or static roadmap planes based on catheter position
Capturing Results Save screenshots and image streams
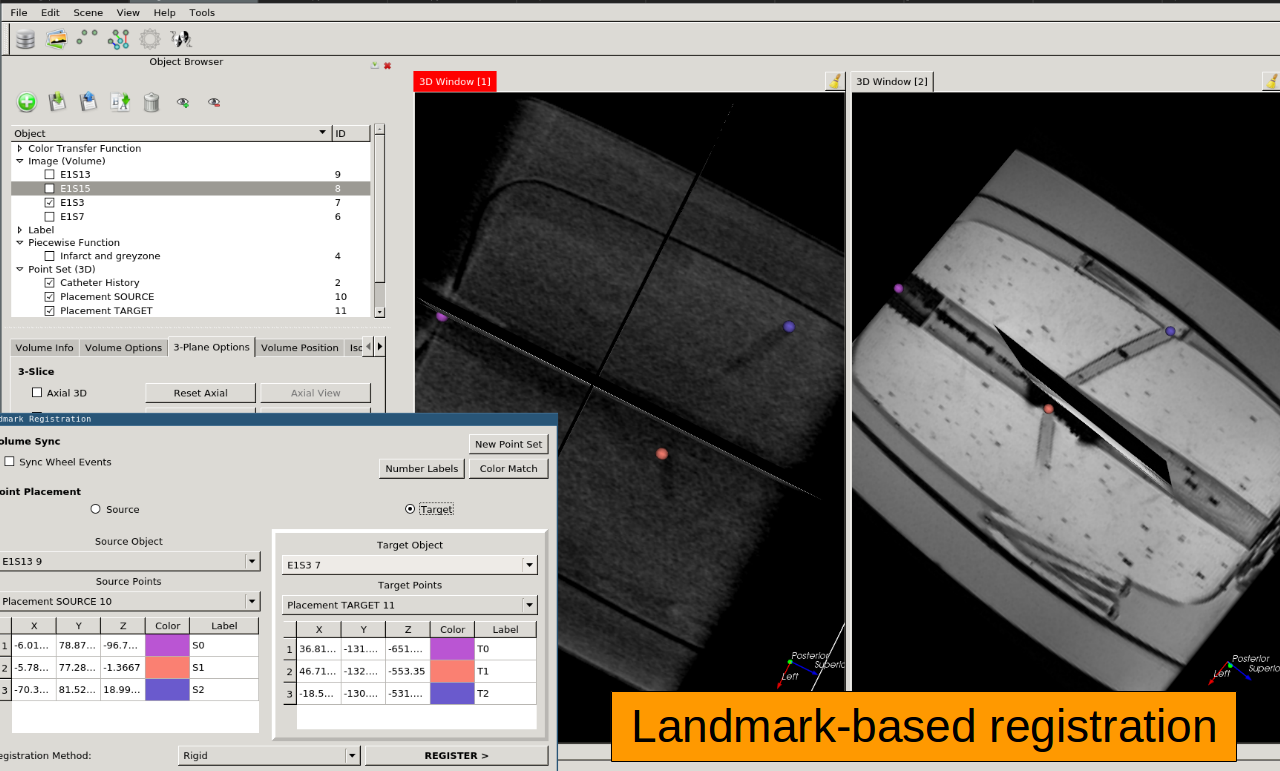
Interactive Landmark Registration Rigid, Affine, Similarity, and Thin Plate Spline landmark-based registration
Extensible Object Framework Define new types of objects for Vurtigo to render
2D Plots Framework for creating and displaying interactive 2D graphs
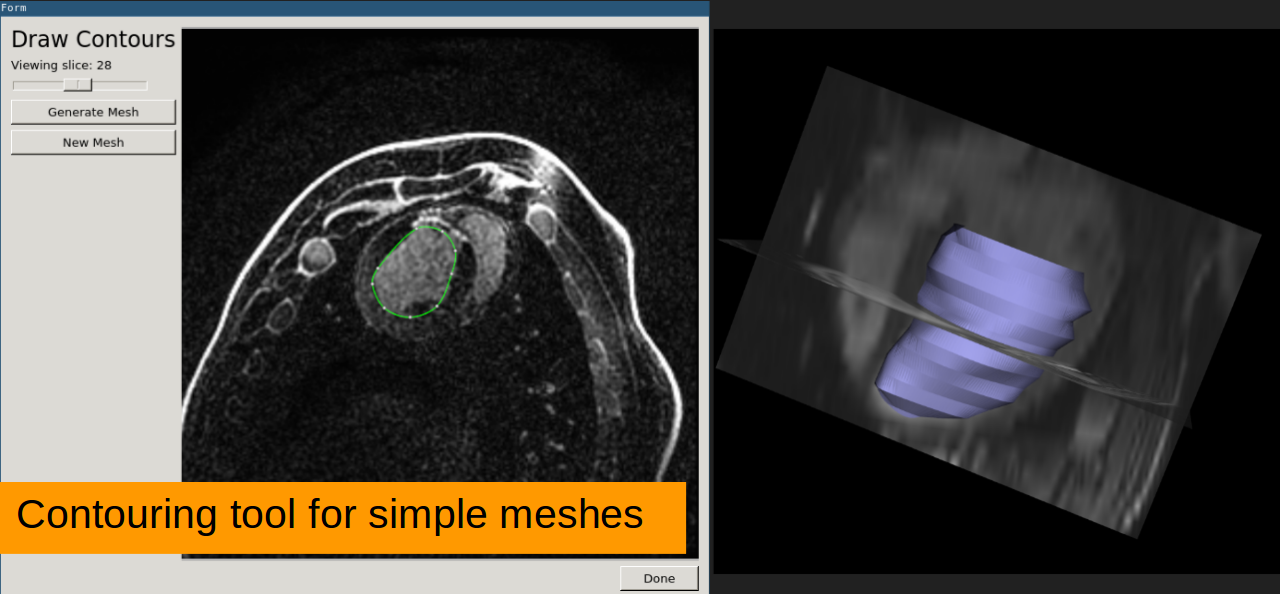
ROI Contours Draw contours on images or surfaces
Mesh tool Create a surface from contours
Mask tool Create a masked volume from contours
Import tool Import images, meshes, and volumes from external formats
 Vurtigo is a four-dimensional (3D + time) real-time visualization software for guiding cardiovascular interventions. It is designed to be part of a pipeline that can connect it to a magnetic resonance imaging (MRI) scanner, actively tracked catheters, and navigational devices.
Vurtigo is a four-dimensional (3D + time) real-time visualization software for guiding cardiovascular interventions. It is designed to be part of a pipeline that can connect it to a magnetic resonance imaging (MRI) scanner, actively tracked catheters, and navigational devices.